The following examples show the performance of three HPC scientific applications on an SGI Altix 3000 system running Linux. These examples are FASTA for bioinformatics, Gaussian for computational chemistry and STAR-CD for computational fluid dynamics. All testing was conducted by SGI.
FASTA Bioinformatics Example
Although biochemistry and computational biology have existed since the mid-1980s, bioinformatics is a fairly new scie.pngic realm, made possible by the convergence of data-intensive life sciences research, laboratory-automation technologies and the development of computer databases and algorithms that can rapidly organize, process and distribute enormous quantities of data. Bioinformatics is used to help bring new drugs to market faster, prevent genetic disease, develop disease- or drought-resistant crops, extend the shelf life of food, find alternatives to petroleum and create foods of the future that will help manage cholesterol or prevent cancer.
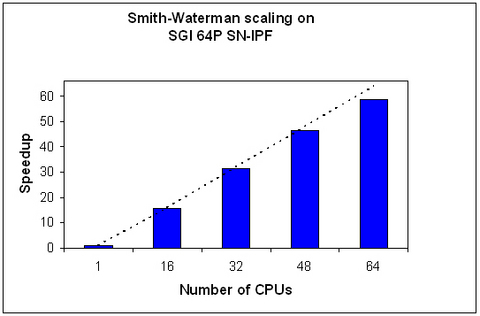
Sequence database searching is among the most critical areas of bioinformatics due to the rapidly growing volume of publicly available biological data. Classical algorithms like Smith-Waterman (T. F. Smith and M. S. Waterman, 1981, Journal of Molecular Biology 147:195-197) provide the most rigorous way to search biological databases for local sequence similarities. However, Smith-Waterman is computationally demanding, so parallelization is extremely important for making this algorithm efficient. One parallel Smith-Waterman implementation available is the FASTA bioinformatics package (W. R. Pearson, 1991, Genomics 11:635-650).
The Smith-Waterman algorithm from FASTA version 3.4 (ssearch34_t), which is available from the University of Virginia (alpha10.bioch.virginia.edu/fasta), was used to measure parallel performance on a 64-processor SGI Altix 3000 system. Smith-Waterman is parallelized with Pthreads, and the SGI ChemBio Applications team also further tuned the core algorithm to more fully exploit the capabilities of the SGI Altix 3000 system. As a result, the Smith-Waterman algorithm shows close to ideal scaling (dotted line represents ideal scaling) with speedups of approximately 59× when running on 64 processors.

National research laboratories, universities, pharmaceutical and biotechnology companies and chemical companies use computational chemistry applications like Gaussian 98 from Gaussian, Inc. (www.gaussian.com) for research in molecular energies, properties and reactions by modeling very large molecular systems using electronic structure methods.
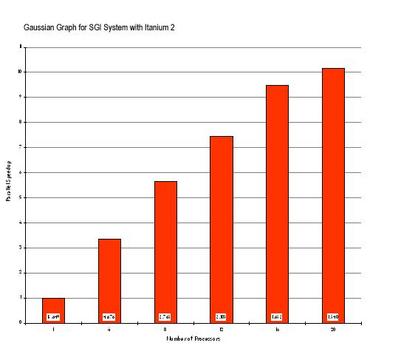
In Figure II, the parallel scaling of Gaussian 98 on a widely used test from Gaussian's QA-suite is shown using an early prototype SGI Altix 3000 system and an earlier version of the Intel compilers targeting Itanium processors rather than Itanium 2 processors. The case corresponds to a Force calculation of the Valynomycin molecule (C54H90N6O18), using Density Functional Theory. The graph shows the parallel speedups for this test case; the labels are the elapsed times in seconds. Even with the early tools and system versions used, the time to perform this calculation is reduced from about 4.5 hours to only 25 minutes when using 20 processors on an SGI Altix 3000 system.
STAR-CD Computational Fluid Dynamics Example
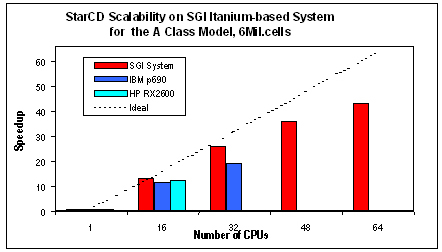
Computational fluid dynamics (CFD) is used in many traditional industries such as automotive, aerospace and power generation. STAR-CD from Computational Dynamics Limited (www.cd-adapco.com) is one of the CFD technology leaders for fluid flow analysis. It can assist a CFD user from initial fundamental design, through parametric studies to optimization, using its advanced physical models and its ability to handle complex geometry using unstructured meshes. STAR-CD runs on all leading UNIX and NT platforms. Its parallel form, STAR-HPC, runs on shared-memory multiprocessor servers, massively parallel systems and clusters of workstations. STAR-HPC uses the MPI library to achieve a high degree of scalability. Using a prerelease version of STAR-CD on an SGI Altix 3000 system with the “Model A Class Dataset” for modeling turbulence flow on an automobile, Linux again demonstrates excellent processor scaling up to 64 processors based on early performance runs. According to published results at www.cd-adapco.com/support/bench/aclass.htm as of early November 2002, Linux even scaled better than two proprietary operating systems: Hewlett-Packard's HP-UX and IBM AIX.